Mol:FL5FAAGS0105
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 43 47 0 0 0 0 0 0 0 0999 V2000 | + | 43 47 0 0 0 0 0 0 0 0999 V2000 |
| − | 1.1286 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1286 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1286 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1286 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8431 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8431 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5575 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5575 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5575 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5575 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8431 5.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8431 5.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4141 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4141 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3004 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3004 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0148 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0148 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0148 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0148 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3004 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3004 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4141 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4141 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7293 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7293 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4438 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4438 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4438 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4438 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7293 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7293 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3004 1.4911 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3004 1.4911 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.1582 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.1582 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0204 2.2718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0204 2.2718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.1582 5.0312 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.1582 5.0312 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7293 1.5099 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7293 1.5099 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1572 -0.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1572 -0.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9822 -0.9415 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9822 -0.9415 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7447 -1.6560 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7447 -1.6560 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1572 -2.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1572 -2.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5562 -2.9230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5562 -2.9230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.3810 -2.9438 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.3810 -2.9438 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7753 -3.6684 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7753 -3.6684 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.3449 -4.3723 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.3449 -4.3723 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5202 -4.3515 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5202 -4.3515 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1259 -3.6268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1259 -3.6268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7393 -5.0969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7393 -5.0969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6724 0.3632 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6724 0.3632 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0736 0.0110 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0736 0.0110 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6104 0.6376 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6104 0.6376 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3915 0.9031 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3915 0.9031 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6455 1.2554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6455 1.2554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1086 0.6289 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1086 0.6289 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3540 1.0208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3540 1.0208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9060 0.9751 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9060 0.9751 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7942 -0.3090 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7942 -0.3090 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3598 -0.2392 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3598 -0.2392 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8442 -0.0144 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8442 -0.0144 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 2 0 0 0 0 | + | 1 2 2 0 0 0 0 |
| − | 2 3 1 0 0 0 0 | + | 2 3 1 0 0 0 0 |
| − | 3 4 2 0 0 0 0 | + | 3 4 2 0 0 0 0 |
| − | 4 5 1 0 0 0 0 | + | 4 5 1 0 0 0 0 |
| − | 5 6 2 0 0 0 0 | + | 5 6 2 0 0 0 0 |
| − | 6 1 1 0 0 0 0 | + | 6 1 1 0 0 0 0 |
| − | 2 7 1 0 0 0 0 | + | 2 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 1 0 0 0 0 | + | 8 9 1 0 0 0 0 |
| − | 9 10 2 0 0 0 0 | + | 9 10 2 0 0 0 0 |
| − | 10 11 1 0 0 0 0 | + | 10 11 1 0 0 0 0 |
| − | 11 12 1 0 0 0 0 | + | 11 12 1 0 0 0 0 |
| − | 12 7 2 0 0 0 0 | + | 12 7 2 0 0 0 0 |
| − | 9 13 1 0 0 0 0 | + | 9 13 1 0 0 0 0 |
| − | 13 14 2 0 0 0 0 | + | 13 14 2 0 0 0 0 |
| − | 14 15 1 0 0 0 0 | + | 14 15 1 0 0 0 0 |
| − | 15 16 2 0 0 0 0 | + | 15 16 2 0 0 0 0 |
| − | 16 10 1 0 0 0 0 | + | 16 10 1 0 0 0 0 |
| − | 14 18 1 0 0 0 0 | + | 14 18 1 0 0 0 0 |
| − | 11 17 2 0 0 0 0 | + | 11 17 2 0 0 0 0 |
| − | 12 19 1 0 0 0 0 | + | 12 19 1 0 0 0 0 |
| − | 5 20 1 0 0 0 0 | + | 5 20 1 0 0 0 0 |
| − | 16 21 1 0 0 0 0 | + | 16 21 1 0 0 0 0 |
| − | 22 23 2 0 0 0 0 | + | 22 23 2 0 0 0 0 |
| − | 22 24 1 0 0 0 0 | + | 22 24 1 0 0 0 0 |
| − | 24 25 2 0 0 0 0 | + | 24 25 2 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 2 0 0 0 0 | + | 26 27 2 0 0 0 0 |
| − | 27 28 1 0 0 0 0 | + | 27 28 1 0 0 0 0 |
| − | 28 29 2 0 0 0 0 | + | 28 29 2 0 0 0 0 |
| − | 29 30 1 0 0 0 0 | + | 29 30 1 0 0 0 0 |
| − | 30 31 2 0 0 0 0 | + | 30 31 2 0 0 0 0 |
| − | 31 26 1 0 0 0 0 | + | 31 26 1 0 0 0 0 |
| − | 29 32 1 0 0 0 0 | + | 29 32 1 0 0 0 0 |
| − | 33 34 1 1 0 0 0 | + | 33 34 1 1 0 0 0 |
| − | 34 35 1 1 0 0 0 | + | 34 35 1 1 0 0 0 |
| − | 36 35 1 1 0 0 0 | + | 36 35 1 1 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 33 1 0 0 0 0 | + | 38 33 1 0 0 0 0 |
| − | 38 39 1 0 0 0 0 | + | 38 39 1 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 33 41 1 0 0 0 0 | + | 33 41 1 0 0 0 0 |
| − | 34 42 1 0 0 0 0 | + | 34 42 1 0 0 0 0 |
| − | 35 43 1 0 0 0 0 | + | 35 43 1 0 0 0 0 |
| − | 36 19 1 0 0 0 0 | + | 36 19 1 0 0 0 0 |
| − | 22 41 1 0 0 0 0 | + | 22 41 1 0 0 0 0 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FAAGS0105 | + | ID FL5FAAGS0105 |
| − | KNApSAcK_ID C00013772 | + | KNApSAcK_ID C00013772 |
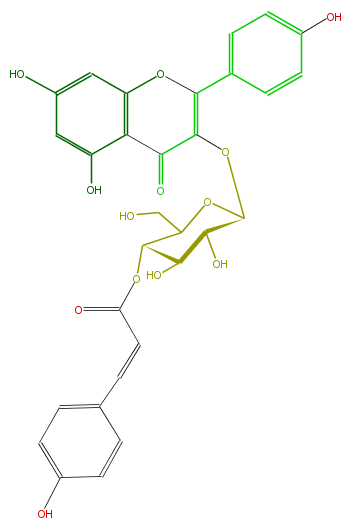
| − | NAME Kaempferol 3-(4''-p-coumaroylglucoside);5,7-Dihydroxy-2-(4-hydroxyphenyl)-3-[[4-O-[3-(4-hydroxyphenyl)-1-oxo-2-propenyl]-beta-D-glucopyranosyl]oxy]-4H-1-benzopyran-4-one | + | NAME Kaempferol 3-(4''-p-coumaroylglucoside);5,7-Dihydroxy-2-(4-hydroxyphenyl)-3-[[4-O-[3-(4-hydroxyphenyl)-1-oxo-2-propenyl]-beta-D-glucopyranosyl]oxy]-4H-1-benzopyran-4-one |
| − | CAS_RN 331448-63-6 | + | CAS_RN 331448-63-6 |
| − | FORMULA C30H26O13 | + | FORMULA C30H26O13 |
| − | EXACTMASS 594.137340918 | + | EXACTMASS 594.137340918 |
| − | AVERAGEMASS 594.51964 | + | AVERAGEMASS 594.51964 |
| − | SMILES OCC(O1)C(OC(C=Cc(c5)ccc(c5)O)=O)C(C(O)C1OC(C3=O)=C(Oc(c4)c3c(O)cc4O)c(c2)ccc(c2)O)O | + | SMILES OCC(O1)C(OC(C=Cc(c5)ccc(c5)O)=O)C(C(O)C1OC(C3=O)=C(Oc(c4)c3c(O)cc4O)c(c2)ccc(c2)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
43 47 0 0 0 0 0 0 0 0999 V2000
1.1286 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1286 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8431 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5575 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5575 4.6844 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8431 5.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.4141 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3004 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.0148 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0148 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3004 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.4141 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7293 3.8594 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4438 3.4469 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4438 2.6219 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7293 2.2094 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3004 1.4911 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.1582 3.8594 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0204 2.2718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.1582 5.0312 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.7293 1.5099 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1572 -0.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9822 -0.9415 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7447 -1.6560 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1572 -2.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5562 -2.9230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.3810 -2.9438 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7753 -3.6684 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.3449 -4.3723 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5202 -4.3515 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1259 -3.6268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7393 -5.0969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6724 0.3632 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0736 0.0110 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6104 0.6376 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3915 0.9031 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6455 1.2554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1086 0.6289 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3540 1.0208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9060 0.9751 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7942 -0.3090 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.3598 -0.2392 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.8442 -0.0144 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 2 0 0 0 0
2 3 1 0 0 0 0
3 4 2 0 0 0 0
4 5 1 0 0 0 0
5 6 2 0 0 0 0
6 1 1 0 0 0 0
2 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 1 0 0 0 0
9 10 2 0 0 0 0
10 11 1 0 0 0 0
11 12 1 0 0 0 0
12 7 2 0 0 0 0
9 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 10 1 0 0 0 0
14 18 1 0 0 0 0
11 17 2 0 0 0 0
12 19 1 0 0 0 0
5 20 1 0 0 0 0
16 21 1 0 0 0 0
22 23 2 0 0 0 0
22 24 1 0 0 0 0
24 25 2 0 0 0 0
25 26 1 0 0 0 0
26 27 2 0 0 0 0
27 28 1 0 0 0 0
28 29 2 0 0 0 0
29 30 1 0 0 0 0
30 31 2 0 0 0 0
31 26 1 0 0 0 0
29 32 1 0 0 0 0
33 34 1 1 0 0 0
34 35 1 1 0 0 0
36 35 1 1 0 0 0
36 37 1 0 0 0 0
37 38 1 0 0 0 0
38 33 1 0 0 0 0
38 39 1 0 0 0 0
39 40 1 0 0 0 0
33 41 1 0 0 0 0
34 42 1 0 0 0 0
35 43 1 0 0 0 0
36 19 1 0 0 0 0
22 41 1 0 0 0 0
S SKP 8
ID FL5FAAGS0105
KNApSAcK_ID C00013772
NAME Kaempferol 3-(4''-p-coumaroylglucoside);5,7-Dihydroxy-2-(4-hydroxyphenyl)-3-[[4-O-[3-(4-hydroxyphenyl)-1-oxo-2-propenyl]-beta-D-glucopyranosyl]oxy]-4H-1-benzopyran-4-one
CAS_RN 331448-63-6
FORMULA C30H26O13
EXACTMASS 594.137340918
AVERAGEMASS 594.51964
SMILES OCC(O1)C(OC(C=Cc(c5)ccc(c5)O)=O)C(C(O)C1OC(C3=O)=C(Oc(c4)c3c(O)cc4O)c(c2)ccc(c2)O)O
M END